A minimization process (least squares fitting) is then executed over a range of -3 to 1 standard deviations about the first guess mean to improve the fit. The standard deviation (SD) or width of the population is then approximated by finding the width of the distribution at 60% of the maximum height. The model initializes by approximating G0/G1 peak as a Gaussian distribution and making an initial guess of the mean by finding the channel with the most cell in the left portion of the data. It assumes only that the data within the G0/G1 and G2/M peaks are normally distributed and that one of those two peaks is identifiable. The Watson model was published by James Watson and colleagues in 1987. It is good laboratory practice to consistently use the same model throughout a study when reporting or publishing statistics. The methods the models employ to calculate their statistics are described below. 
Consequently, results from one model may vary quite significantly from the other. The two models differ in their mathematical calculations of each phase of the cell cycle. The univariate model will appear by default as shown in Figure 1.įlowJo v10 provides two univariate cell cycle platforms, the Watson Pragmatic algorithm (1) and Dean Jett Fox (DJF) (2). To launch the univariate cell cycle model click on the population of interest in the workspace, then select the Cell Cycle task from the Biology Band. This signifies that option-clicking a layer in the legend of a graph will delete that layer from the graph.Univariate modeling can be used to create a fit to cell cycle data based on statistics in one dimension, traditionally DNA content.įlowJo provides a simple interface to performing fairly sophisticated DNA/Cell Cycle analysis. If you hold down the option key, you'll see that the cursor changes from a vertical reordering tool into a trash can. Dragging the order of the layers will cause the graph to be redrawn, so you can quickly see which layers you want to feature in the top positions. 
Because events in front can obscure others, you may want to have the smaller populations on top of the larger ones, though this depends greatly on the context of your analysis. Moving a graph's name to the top of thelist moves the graph to the top of the overlay stack. The order of the lines in the legend mirrors the order of the graphs displayed. 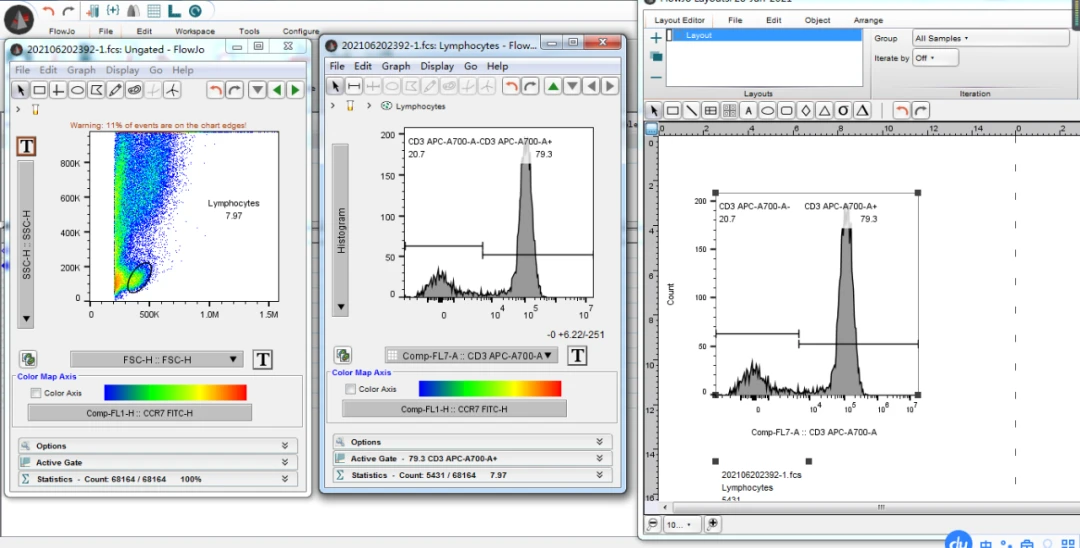
This tools allows you to reorder the legend. If you move the mouse over the text description of the layers, you'll see the cursor become a pair of bars with vertical arrows sticking out of it. Clicking the mouse on the outside edge of the legend lets you move or resize the legend. When the mouse passes over an edge of the legend, the cursor will become a hand. Drop the second graph (or more, if you had multiple nodes selected in the workspace) on top of the original, and an overlay is automatically created. As the mouse moves into the original graph item, you'll see that it highlights its borders to signify that it will accept the contents of the drag. Then drag a second on top of the first one. To create an overlay, drag a graph into the Layout Editor. Once an overlay has been defined, the layout editor can create the same graph for many different samples or sets of samples, by iterating over a group in the workspace. Within the Layout Editor you can edit the color and order of the items, to customize the look of the overlay graph that best highlights your data. Overlays may be created on either univariate (right) or bivariate displays (below). This makes it easy to compare the data, using differing colors to show multiple populations at the same time. Often it is useful to plot two or more data sets or population on the same axes.
0 Comments
Leave a Reply. |
AuthorWrite something about yourself. No need to be fancy, just an overview. ArchivesCategories |
 RSS Feed
RSS Feed
